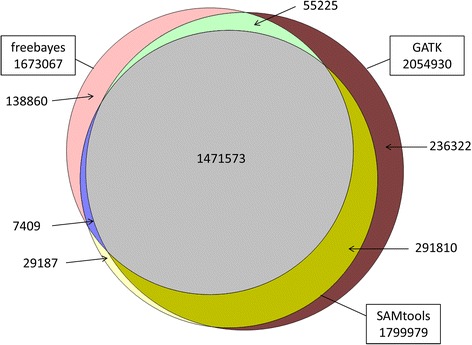
Comparison among three variant callers and assessment of the accuracy of imputation from SNP array data to whole-genome sequence level in chicken.
BACKGROUNDThe technical progress in the final decade has made it attainable to sequence tens of millions of DNA reads in a comparatively quick time-frame. Several variant callers primarily based on totally different algorithms have emerged and have made it attainable to...
